- Research
- Open access
- Published:
Role of CNTNAP2 in autism manifestation outlines the regulation of signaling between neurons at the synapse
Egyptian Journal of Medical Human Genetics volume 22, Article number: 22 (2021)
Abstract
Background
Autism is characterized by high heritability and a complex genetic mutational landscape with restricted social behavior and impaired social communication. Whole-exome sequencing is a reliable tool to pinpoint variants for unraveling the disease pathophysiology. The present meta-analysis was performed using 222 whole-exome sequences deposited by Simons Simplex Collection (SSC) at the European Nucleotide Archive. This sample cohort was used to identify causal mutations in autism-specific genes to create a mutational landscape focusing on the CNTNAP2 gene.
Results
The authors account for the identification of 15 high confidence genes with 24 variants for autism with Simons Foundation Autism Research Initiative (SFARI) gene scoring. These genes encompass critical autism pathways such as neuron development, synapse complexity, cytoskeleton, and microtubule activation. Among these 15 genes, overlapping variants were present across multiple samples: KMT2C in 167 cases, CNTNAP2 in 192 samples, CACNA1C in 152 cases, and SHANK3 in 124 cases. Pathway analysis identifies clustering and interplay of autism genes—WDFY3, SHANK2, CNTNAP2, HOMER1, SYNGAP1, and ANK2 with CNTNAP2. These genes coincide across autism-relevant pathways, namely abnormal social behavior and intellectual and cognitive impairment. Based on multiple layers of selection criteria, CNTNAP2 was chosen as the master gene for the study. It is an essential gene for autism with speech-language delays, a typical phenotype in most cases under study. It showcases nine variants across multiple samples with one damaging variant, T589P, with a GERP rank score range of 0.065–0.95. This unique variant was present across 86.5% of the samples impairing the epithelial growth factor (EGF) domain. Established microRNA (miRNA) genes hsa-mir-548aq and hsa-mir-548f were mutated within the CNTNAP2 region, adding to the severity. The mutated protein showed reduced stability by 0.25, increased solvent accessibility by 9%, and reduced depth by 0.2, which rendered the protein non-functional. Secondary physical interactors of CNTNAP2 through CNTN2 proteins were mutated in the samples, further intensifying the severity.
Conclusion
CNTNAP2 has been identified as a master gene in autism manifestation responsible for speech-language delay by impairing the EGF protein domain and downstream cascade. The decrease in EGF is correlated with vital autism symptoms, especially language disabilities.
Background
Autism is a neurodevelopmental condition with high heritability and a complex genetic mutational landscape. It has been characterized by social communications deficits, restricted interactions, and repetitive behavior patterns and interests [1]. Its prevalence rate is 1 in 59 children worldwide [2]. Landmark symptoms for autism include hypersensitivity, impulsivity, agitation, mood swings, and mild to severe cognitive functions impairment [3]. These symptoms range from above-average to intellectual disability, accompanied by seizures and language impairment. Defective cross-functionality in relevant domains and other cranial defects in subjects results in autism manifestation, generally before the age of three [4]. Speech-language delay is a unique and empirical phenomenon observed in autistic children. It is crucial to study the causality and molecular markers involved with autism with speech-language delays [5].
Unequivocal genes causative for autism pathophysiology has not been pinpointed, even after decades of autism research advancements starting from linkage to next-generation sequencing techniques to date. Attempts focused on linkage and candidate studies to understand autism-specific variants have implicated several significant findings [6]. Genome-wide association studies (GWAS) were scalable to neuronal function and corticogenesis, which provided confidence to identify risk autism variants viz. PTBP2, CADPS, and KMT2E [7]. However, GWAS could not confirm the detection of strong contributors to common alleles for autism. Association between autism and specific Mendelian disorders has been observed; for example, PTEN macrocephaly is associated with autism severity [8]. Specific copy number loci have been associated with autism with statistical significance values. Various microRNA recognition elements (MRE) modulating single-nucleotide polymorphisms (SNPs) and MRE-creating SNPs present in the 3′ UTR of autism have significant implications. These genes have a notable effect on autism manifestation and severity [9]. Limited information obtained from GWAS, genotyping, and other processes has directed the interest of researchers towards rare variants and point mutational studies. Autism-associated point mutations in various genes have added information and clarity to its molecular basis for manifestation. De novo and other types of point mutations have been identified in 15–20% of the autism subjects [10].
Whole-exome sequencing (WES) can be used as a reliable tool to scan various exomes and identify causal variants for autism. WES helps explore the exome for rare autism-specific de novo and transmitted variants disrupting proteins [11]. It can help to evaluate whether the co-occurrence of de novo events in the same individual increases risks for autism or not. These sequence datasets showcase over 120 casual genes with clinically relevant genetic variants identified for autism in the last decade [12]. These variations can be point mutations, insertions, deletions, and copy number variations in the coding regions, in either the homozygous or the heterozygous state [13]. Multiple levels of sequencing advancements have opened new, quicker, and cost-effective avenues to date. Several SNPs with recurrent deleterious mutations in ARID1B, SCN1A, SCN2A, and SETD2 genes have been reported so far. These identified mutations result in gain or loss of function of one or more functional copies of a gene or contiguous genes besides biallelic mutations of both gene copies, which is suggestive of contribution to autism susceptibility [14]. Numerous genes have been well established with validation in several cohorts to elucidate gene dosage sensitivity and relevance to autism-specific pathways [15]. Although numerous studies have been conducted on autism using the whole-exome sequencing technique yet, its pathophysiology is not thoroughly addressed.
Disease susceptibility for heterozygous variants is influenced by the analysis of the haploinsufficiency and probability of being the loss-of-function–intolerant (pLI) scores for mutated genes [16, 17]. It reflects the intolerance to loss-of-function and deleterious mutations via the functional impact of essential genes (EGs) commonly observed in heterozygous mutations. These scores indicate the cumulative effect of deleterious variants in EGs on complex neurodevelopmental disorders such as autism with a threshold of > 20 [17]. Therefore, this study aims to identify high confidence genes for autism and study their interplay with an in-depth analysis of one master gene with gene variants and associated causative pathways. It pinpoints damaging heterozygous variants in CNTNAP2, a significant gene in autism, and delayed speech-language phenotype.
Methods
The present study consists of 222 whole-exome sequences deposited by Simons Simplex Collection (SSC) at the European Nucleotide Archive with the accession number PRJNA167318. The SSC group has carried out the sequencing using Illumina Genome Analyzer IIx paired-end sequencing platform at the coverage of 100X, with library preparation according to Illumina protocols. In the current study, we performed an exhaustive analysis to identify the high confidence autism genes in the 222 exome sequences of quartet sample sets from the SSC family. Each family under study comprises an autistic subject with unaffected siblings and parents with a detailed case history. Only the affected probands have been studied under the current investigation.
Whole-exome sequences in .fastq format were aligned against the hg19 build of the human reference genome using the Strand Next-Generation Sequencing (NGS) platform due to its accuracy, correctly mapped reads, and receiver operating curves. Post-alignment quality check was performed to remove false-positive variant reads. The sequence data was run on multiple platforms: STRAND NGS, Partek, and direct command prompt algorithm-based pipelines to avoid further false positives. Partek® software (©Partek Inc., St. Louis, MO, USA) and Strand NGS software (Version 2.8, Build 230243 ©Strand Life Sciences, Bangalore, India) were used for the analysis along with direct command prompt algorithms softwares: Burrows-Wheeler Aligner (BWA)-backtracks [18] and Bowtie 2 [19]. BWA and Bowtie 2 have been known as ultrafast and memory-efficient tools for mapping human exome or genomes. Comparative results were obtained at pre-and post-alignment, along with multi-level quality checks (QCs). All the variants identified across multiple platforms were considered for the study and exported in variant calling format (vcf).
Further, variants were called using variant calling program web ANNOVAR (wANNOVAR) with vcf files as input. wANNOVAR software was used to annotate the variant files based on position, gene, amino acid change, zygosity, and mutation effects. Variant calls with a read depth of ≥ 20 were included in the study. Minor allele frequency was limited to P value ≤ 0.05 based on the EXAC and 1000G studies. The candidate genes were filtered for deleterious and damaging mutations—stop gain, stop loss, missense, and nonsynonymous. Pathogenicity scores were calculated across eleven platforms with a minimum threshold of a mutation being tagged as damaging across at least five platforms to be considered for further analysis. The priority-based classification for the known autism candidate genes was performed using the Simons Foundation Autism Research Initiative (SFARI) gene list [20]. PredictSNP tool was used to predict the effects of identified mutations on protein function for prioritization for further characterization using six different robust prediction classifiers for disease-related mutations.
Haploinsufficiency and pLI scores for heterozygous mutations have been used to support the clinical interpretation of novel loss-of-function variants with gene prioritization for whole-exome sequencing [21]. It is calculated to understand and validate the presence of heterozygous mutations and their functionality [16]. The pLI scores indicate the tolerance level of a given gene to loss of function (LoF) based on the number of protein-truncating variants. Thus, the stop gains and frameshift variants are referenced in the human genome using gene size and sequencing coverage metrics. It is often used for the prioritization of candidate genes. For LoF mutations, assumptions are set for three gene classes: null (where LoF variation is completely tolerated), recessive (where heterozygous LoFs are tolerated), and haploinsufficient (where heterozygous LoFs are not tolerated) for tolerance to LoF variation. Observed and expected variant counts have been used to determine the probability that a given gene is extremely intolerant of LoF variation. The closer pLI is to one, the more the gene is intolerant to LoF. pLI ≥ 0.9 is considered an extremely LoF-intolerant gene set [22].
Enrichment analysis for the identified genes was performed through KEGG pathways and an extensive literature review. Gene-enriched pathways relevant to autism were selected along with other known associated genes. BIOSTRING software was used to create gene-gene, gene-protein, and protein-protein interaction networks using the top ten genes.
The disease pathway was created using Ingenuity Pathway Analysis (IPA) with an inbuilt database of curated literature. Upstream and downstream of the mutant genes were overlaid. The pathway enrichment was performed using Z-score, P value, and Jaccard similarity testing to identify enriched disease pathways with disruptions/blocks caused due to mutations [23,24,25,26].
Protein modeling was performed using STRUM software [27]. STRUM is a method for predicting fold stability change (ΔΔG) of protein molecules upon single-point nonsynonymous mutations. It adopts a gradient boosting regression approach to train the Gibbs free-energy changes on various features with different levels of sequence and structure properties [27]. Its uniqueness lies in combining sequence profiles with low-resolution protein structure models from structural prediction. This process enhances the method’s robustness and accuracy, making it applicable to various protein sequences, including those without experimental structures. It starts from wild-type sequences and constructs 3D models by the iterative threading assembly refinement simulations. DOMPred tool was used to derive a graph from the aligned termini positions using PSI-BLAST local alignments. In this case, larger values indicate regions with sequence discontinuities in putative domain boundaries. This also gives the predicted number of domains and the positions of domain boundaries for the predictive peaks. The graph can be visualized to confirm the predicted number of domains and possible domain boundaries. In case of a mutated protein, a larger degree of variation is possible due to disorder and variation in the domain linker region aspects [28].
Results
The authors performed WES analysis for 222 autism subjects with 100X coverage and a 95% confidence interval. The mean read length of ≥ 100bp with 10.19 GB of raw data was obtained from the sequencing reaction. It generated a total of 47 million reads with read quality in terms of a Phred score of 33.32 forward read (R1) and 33.10 reverse read (R2). The Quality scores were 27.65% and 66.47% for R1 and R2, respectively (Fig. 1). The quality check for pre-and post-alignment was of an appropriate standard. Alignment breakdown was marked at ≥ 90% with unique paired alignment in almost all the cases, while the unaligned part was minimal. Local alignment captured 10,000 variants on an average in the SNP processing step before SNP detection was conducted with false discovery rate (FDR) set at ≤ 0.5.
The identified 10,000 variants were annotated across the regulatory, untranslated regions, exons, introns, downstream, and intergenic regions using SNP detection. On analysis of the exonic variants from the annotated whole-exome sequencing dataset, 943 genes were identified as damaging for autism. On applying SFARI gene scoring, identified genes with scores 1 and 2 were 192 and 182 in number, respectively. Further, on filtering the genes based on pathogenicity and haploinsufficiency scores, the gene list was streamlined to 15 genes with 24 variants. Annotation of these genes revealed rare and deleterious variants for KMT2C, CNTNAP2, CACNA1C, SHANK3, ANK2, HECTD4, MAP1A, SKI, SCRAP, CUL7, ZNF804A, CNTNAP3, CACNA1H, LRP1, and CNTN4 genes (Table 1). These variants belonged to either nonsynonymous, frameshift insertions-deletions, or stop gain variants in the coding regions. This mechanistic study was assisted by exhaustive literature and SFARI gene scoring for in silico validation. The burden of gene variants was observed on chromosomes 1, 2, 3, 4, 6, 7, 11, 12, 14, 16, and 22 (Table 1).
Among these 15 genes, the study of overlapping variants presents across all the samples revealed exclusive variants for KMT2C in 167 cases, CNTNAP2 in 192 samples, CACNA1C in 152 cases, and SHANK3 in 124 cases (Table 1). Previously reported variants were identified for the filtered gene sets for the reported mutational landscape.
Pathway analysis of the 15 high confidence genes was outlined clustering of autism-relevant genes WDFY3, SHANK2, CNTNAP2, HOMER1, SYNGAP1, and ANK2 with several primary and secondary physical interactors. These genes are encircled and highlighted in red in the schematic pathway represented in Fig. 2. Autism-related processes and phenotypes were enriched in the pathway obtained.
Pathway analysis for the high-risk autism genes using Ingenuity Pathway Analysis (IPA). Clustered protein within the top network-associated genes as derived from IPA algorithms is shown. Proteins identified are encircled and labeled with the protein symbol in red fill. Direct connections between/among proteins are shown in solid lines; indirect interactions are shown as dashed lines or edges. The constructed pathways analysis of the 15 high-risk genes has shown an association with various autism phenotypes and processes, including learning, synaptic transmission, abnormal social behavior, and social withdrawal. These genes coincide with multiple pathways with overlapping connections across different pathways and present in the upstream and downstream of vital processes. Each gene has a divergent and convergent pathway and can pave a path towards autism manifestation
CNTNAP2 showed a haploinsufficiency score of 4.94 with nine damaging nonsynonymous variants—T589P, T118P, H764P, G285A, A588P, W134G, N139S, N139S, R160H, and T831S in seven exons with GERP rank score range of 0.065–0.95. CNTNAP2 had shown exclusive variants with relevant read depth and P value in 86.5% of the cases under investigation (Table 2). Based on the stepwise analysis and interpretations, CNTNAP2 was selected for the downstream analysis. Minor allele frequency ranges from 1.65 × 10−05 to 0.216, with the highest read depth being 210. One unique variant was identified in 192 cases positioned at 589, resulting in the amino acid change from threonine to proline, impairing the epithelial growth factor (EGF) domain with a mutation-induced perturbation of protein folding stability change, ΔΔG value of 2.67 kcal/mol (Fig. 3). The amino acid change T589P was present across 91.06% of the sample cohort. It has an overall confidence score of 87% with deleterious effect across multiple pathogenicity platforms, calculated using the PredictSNP tool. Established miRNA genes, miRNA548AQ, and miRNA548F were mutated within the CNTNAP2 region, adding to the severity in the cohort.
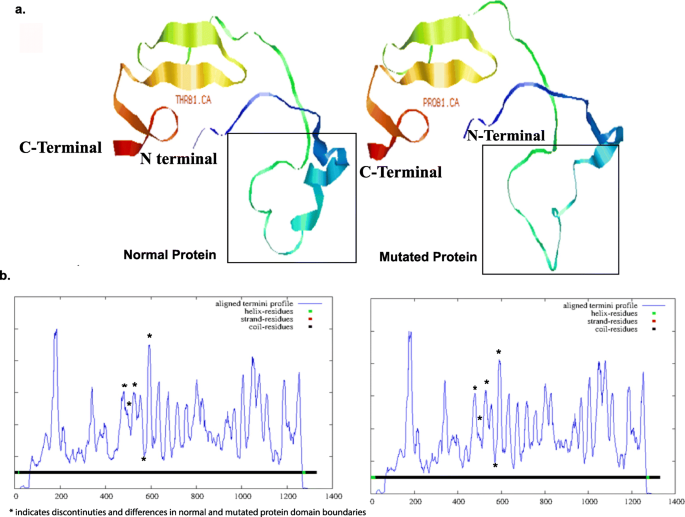
The protein structures for the normal and the mutated CNTNAP2 protein were modeled by STRUM using the CNTNAP2 protein structure with PDB ID 5Y4M as a template. The variants showed a range of ΔΔG ≥ 0.5, indicating stability in mutational sensitivity. The protein folding was distorted and disoriented in the mutated protein. The unique variant T589P, present across multiple populations, showed reduced stability by 0.25, increased solvent accessibility by 9%, and reduced depth by 0.2 in mutant protein product (Fig. 4). The secondary structure underwent negligible changes across normal and mutated protein structures. Ten domain boundaries positioned at 185, 341, 481, 594, 677, 802, 967, 1048, and 1253 were predicted for the modeled normal protein using the DOMPred tool (Marsden, R). These have undergone changes positioned at 174, 341, 478, 591, 717, 802, 967, 1078, and 1253 in terms of coiling and aligned termini profile in modeled mutated protein (Fig. 4). Residues in terms of helix, coiling, and strand have been indicated with a center at the 700th residue from the domain boundaries.
a Protein modeling for normal and mutated CNTNAP2 protein product with marked mutation and conformational change highlighted in a box. The mutated protein structure was rendered as non-functional with reduced stability and depth and increased solvent accessibility. b Domain boundary prediction to understand the changes in residues and aligned termini profile. The coil residue have undergone multiple changes highlighted with an asterisk in the graph across normal and mutated protein leading to disorder and variation
Network analysis of CNTNAP2 protein revealed various physical interactors: CNTN2, CALM1, CALM3, CACNB1, CACNB2, ANK2, ZNF804A, CACNA1H, and CACNA1C proteins with varying interaction scores between 0.5 and 1. Interestingly, all the interactors enriched have also been mutated in the dataset under study contributing to the manifestation of autism through different entry-exit points. The average local clustering coefficient was significant at 0.548, with a P value of 0.00103. The study of each physical interactor’s expression levels revealed shared expression patterns of CNTNAP2 with CACNB2, CALM1, CNTN2, and CALM3 proteins in the human model (Supplementary Figure 2).
Discussion
Deciphering causal genes in autism has been difficult due to its genetic heterogeneity, varied pathogenicity, and associated comorbidities. WES is employed as a single genetic tool with high-throughput computational program to identify causal gene variants and disease pathways for autism [26]. The confidence interval of 95% with an optimal insert length of 200 bp indicates effective enrichment, leading to sequencing results. Appropriate coverage of 100X with a significant Phred score and read depth enhances the confidence with 98% sensitivity and higher positive predictive values for nonsynonymous variants [45]. Quality check scores and mean alignment breakdown values are typical and follow the sequenced data’s default value range. Significance and confidence of detected rare variants depend on the sequence quality, sample size, and the prior probability that the allele exists. The difference in coverage range for variations is 15X across the sample cohort—optimal according to standard sequence data protocol [46]. FDR rate below 0.5 directs towards increased confidence for the variants to be crucial for disease manifestation.
For SNP detection and identifying genes, rare coding variants are selected with a P value of ≤ 0.05 for stringent and accurate correlation of variants to autism. The universally accepted Gene Score database housed at SFARI places each gene into a category with a score based on relevant evidence available. These scores indicate the gene’s relevance and severity in causing autism [47]. Identified variations belong to regulatory regions, exons, introns, and downstream regions in protein-coding and non-protein-coding regions. Multiple genes containing damaging variations have been observed across all samples. Utilization of a custom-developed pipeline with stringent filters revealed 24 significant disease-causing variants. Coding variants are taken into consideration for the downstream analysis focusing on deleterious/damaging variants. There are multiple evidence lines to direct the cumulative effect of deleterious/damaging coding variants for autism [48]. Out of the 15 identified genes, six gene variants with high P value and damaging variants have been shown to disrupt the normal gene function adversely. The identified chromosome burden complies with the previous trends for autism. Each gene identified plays a crucial role in autism manifestation in a unique, well-defined manner [49].
KMT2C, CNTNAP2, and SHANK3 are well-established causal genes for autism coupled with speech-language disabilities and delays [29, 50, 51]. These genes showed damaging mutations impairing crucial protein domains related to speech-language processes and autism. A phenotype-genotype correlation could be set for these genes for speech-language disability, a crucial phenotype in 90% of the sample cohort. For instance, several variants in SHANK3 with a non-functional SRC Homology 3 domain are known to impair the dendritic spine morphology and synaptic transmission, resulting in autism with delayed speech-language [52, 53]. Similarly, KMT2C and CACNA1C have established trajectories for autism manifestation [30, 54]. Despite convergent evidence from multiple studies, the CNTNAP2 gene shows the strongest association with autism [55]. Knockout mice experiments with CNTNAP2 show striking similarities with autism symptoms [56]. CNTNAP2 gene is well established, yet connections remain to be explored, making it an exciting gene to study further. CNTNAP2 is well established
In parallel, the pathway enrichment analysis constructs gene clusters, which showed association with various autism phenotypes and processes such as learning, synaptic transmission, abnormal social behavior, and social withdrawal. Previous studies in autism using machine learning have reported 77 such gene clusters with significant enrichment in crucial pathways in autism pathophysiology, ultimately resulting in autism manifestation [57]. They have overlapping connections across different pathways and present in the upstream and downstream of vital processes. Each gene has a divergent and convergent pathway and can pave a path towards autism manifestation, as shown by multiple study groups [49, 58, 59].
Sequential criteria based on filtration of genes and overlapping of genes in various studies; based on the parameters of haploinsufficiency and pLI, the prevalence of variants, pathogenicity, and gene selection criteria, and the pathway clustering, CNTNAP2 was identified as a high-risk autism gene. Considering overlapping studies using multi-facet criteria of damaging variants and impaired pathways, CNTNAP2 has shown damaging/deleterious, nonsynonymous, stop gain, and frameshift deletion variants with a haploinsufficiency score of ≥ 4.94 calculated at a read depth (RD) of > 70. A connecting link for CNTNAP2 and autism was established through its biological functionality. CNTNAP2 is present in the synaptic junction impairing axonal growth at cortisol neurons [60], responsible for language ability-vital to autism [58].
CNTNAP2 is the crucial player in synaptic plasticity, localized at myelinated axons associated with potassium channels. It functions in the nervous system of vertebrates as cell adhesion molecules and receptors. The chromosome position of 2.4-Mb-sized CNTNAP2 is 7q35-q36.1, which comprises of 24 exons in total. The gene variant identified in 192 cases lies in exon 11 (Table 2, Fig. 3). CNTNAP2 encodes CAM that regulates signaling between neurons, highly expressed in neurons that control language and language development difficulties. It encodes CASPR2 with expression restricted to neurons (transmembrane scaffolding protein) clustering voltage-gated potassium channels at the Nodes of Ranvier. It plays a significant role in “language development” in autism. It is highly expressed in a cortical–striatal–thalamic circuit, involved in diverse higher-order cognitive functions.
Interestingly, previous studies have identified SNP clusters in intronic regions to be associated with communicative behavioral delays in screening normal healthy cohorts. Genetic variance at this locus is suggestive of its role in language endophenotypes [61]. Associations of CNTNAP2 have been identified with crucial autism phenotypes and speech-language impairment [29].
CNTNAP2 showed nine damaging variants with relevant GERP scores and appropriate read depth. Considering the threshold values of GERP, PolyPhen, and SIFT pathogenicity scores enhanced the probability of the selected variants to be disease-causing [62]. Eight out of the nine variants have been cataloged as causal variants with read sequence identifiers and the associated protein domains and motifs impaired. ΔΔG value could be used to predict the variant’s effect on protein folding and perturbations caused by it. A value of ≥ 0.5 kcal/mol is considered destabilizing, enhancing the effect of dysfunctional protein on disease severity [63].
The deleterious nature and high confidence of the overlapping CNTNAP2 variant were considered for the downstream analysis. It is present in the protein coiling of the protein and present across 192 samples. It covers the epithelial growth factor (EGF) protein domain, which is involved in the proliferation and differentiation of nervous tissue during neurogenesis and promotes wound healing [64]. SNPs for EGF play a significant role in the etiology of abnormal behavior in children with autism. Decreased EGF plasma levels are correlated with hyperactivity, decreased motor skills, and the tendency for tiptoeing [64]. The decreased EGF could be due to the increased ligand binding to its receptor resulting in increased EGFR and decreased EGF. This suggests their association with the etiology of autism [64]. Gene disruptions can affect the CNTNAP2 expression through regulatory miRNAs through deletion or duplication in miRNAs. miRNAs affect the cell differentiation in neuronal cells by downregulation of non-sense mediated RNA decay of genes involved in neurodevelopment [65]. Among its potential targets are a few of the notable autism genes—PTEN, SLC1A1, GRIK2, GABRG1, and GABRA4, which have been evaluated through miRNA expression profiling of cell-derived total RNA [66]. For CNTNAP2, ten miRNAs have been identified in regulatory networks as a hub with transcription factors coupled with target genes. However, no such associations have been established so far for CNTNAP2 expression regulatory miRNAs. Intron 3 of CNTNAP2 shows the deletion of one copy of miR548AQ and miR548F in several patients of autism, as evident from the current investigation as well [67].
Protein modeling shows conformational changes in the structure of the mutated protein product. The normal protein, on having a mutation at 589, undergoes a change in ligand to protein binding by introducing a premature stop codon that is predicted to produce a non-functional, impairing secondary structure. The stability is considerably reduced in the mutated proteins in the current investigation indicating a more considerable impact on the protein folding and disease manifestation. The change in Gibb’s free energy upon protein folding increase in negative value is indicative of greater stability. Substitution in protein sequence due to the presence of an SNP can result in a change in ΔΔG, wherein 0.5 and 0.5 kcal/mol indicates stabilizing and destabilizing mutations, respectively [68]. Increased solvent accessibility would expose much more residual active sites of the protein structure, resulting in easier replacement of amino acid for conformation in secondary structure [69, 70]. Destabilizing mutational protein folding renders disease susceptibility, which has a much more collective effect than the stabilizing counterparts. The mutant protein product would contain only extracellular domains secreted from the cell, which can further have additional deletions resulting in impaired protein [61]. Due to random coiling at the predicted domain boundaries, it would result in a disordered protein structure with fewer structural elements with the vast majority of residues’ solvent exposed [71]. This would reduce the protein non-functional with discontinuous domains. Domain boundary analysis is crucial to understand the local decrease of protein structural constraints, with variants present [72]. Also, it reduces specificity, which would ultimately decrease the sensitivity in the ROC curve [28, 73]. This would help understand the protein folding in secondary structure stability and other properties for autism-related functionality. Such studies should be warranted for protein level studies in autism in larger cohorts.
Rare variants in the CNTNAP2 gene, including deletions and nonsynonymous changes, have indicative roles in autism, intellectual disability, developmental delay, and language impairment [29]. Autism shows genetic heterogeneity, and hence, convergence and divergence of gene clusters are often seen in previous studies. Physical interactors of CNTNAP2 protein-containing variants in the sample cohort aggravate the mutational sensitivity and affect the more significant impact. Taking these interactions into account, a model can be put forth with CNTNAP2 protein as a bridge to connect different cell type variants through CNTN2. Disruptions in such bridges are vital to accounting for the various genotypes, and phenotypes accounted for intracellular variations of CNTNAP2 and the intact and mutant protein roles using model animals [74]. Several studies provide genetic evidence for the CNTNAP2 gene to be closely related to altered gene expression in autism brain [56]. CNTNAP2 directly interacts with FOXP2 suggesting a link between language impairment and autism circuital pathway [75]. Mutations in CACNA1H and CACNA1C, which form the alpha subunits and CACNB1 and CACNB2 of the subunits, impair the multiprotein complex calcium voltage-gated channels.
Similarly, various gradients are established and maintained by a rich array of calcium pumps, exchangers such as ANK2, phosphorylases CALM1 and CALM2, voltage-activated, and ion channels, with an array of calcium-binding proteins. These permit tight regulation of calcium concentrations in cytosol and intercellular spaces and downstream signals. These functionalities are essential for normal cognitive functions, especially synaptic plasticity, memory, the excitability of neurons, neurotransmitter emission, axon growth, and neurons [76]. Adhesion molecule CNTN2 and cell recognition molecule CNTNAP2 aid the process in downstream analysis.
Therefore, rare mutations in CNTNAP2 present in the EGF domain could be critical players in the manifestation of autism along with its physical interactors. The gene could act as a marker to warrant studies at the molecular level for further insights into the underlying pathways.
Conclusion
Autism exhibits genetic heterogeneity, and hence, it becomes difficult to pinpoint one single gene for its manifestation. The gene clusters with varied pathways show the convergence of multiple gene variants, resulting in autism manifestation. Whole-exome sequencing proves to be a reliable tool for deciphering the causal genes for autism manifestation. Deciphering the autism exome identified the mutational landscape derived from single and multi-base DNA variants. Genes carrying mutations were identified in synaptogenesis processes, EGF signaling, and PI3K/MAPK signaling. Protein-protein interactions of NrCAM and CNTN4 with CNTNAP2 increased the impact and burden on autism.
Limitation of the study
A detailed study in a larger cohort with parental and sibling exome analysis could be warranted to identify familial markers. Overlapping studies could be performed on similar datasets re-sequencing techniques. Further, the variants identified could be validated using Sanger sequencing if samples were available.
Availability of data and materials
The sample cohort consisting of 222 whole-exome sequences for the present study has been deposited by Simons Simplex Collection (SSC) at the European Nucleotide Archive under the accession number PRJNA167318.
Abbreviations
- BWA:
-
Burrows-Wheeler Aligner
- EGF:
-
Epithelial growth factor
- EGs:
-
Essential genes
- FDR:
-
False discovery rate
- GWAS:
-
Genome-wide association studies
- IPA:
-
Ingenuity Pathway Analysis
- LoF:
-
Loss of function
- MicroRNA:
-
miRNA
- MRE:
-
miRNA recognition elements
- NGS:
-
Next-Generation Sequencing
- pLI:
-
Probability of being loss-of-function intolerant
- QC:
-
Quality check
- R1:
-
Forward read
- R2:
-
Reverse read
- SFARI:
-
Simons Foundation Autism Research Initiative
- SNP:
-
Single-nucleotide polymorphism
- SSC:
-
Simons Simplex Collection
- vcf:
-
Variant calling format
- wANNOVAR:
-
Web ANNOVAR
- WES:
-
Whole-exome sequencing
- ΔΔG:
-
Fold stability change
References
Eapen V, McPherson S, Karlov L, Nicholls L, Črnčec R, Mulligan A (2019) Social communication deficits and restricted repetitive behavior symptoms in tourette syndrome. Neuropsychiatr Dis Treat. https://doi.org/10.2147/NDT.S210227
Baio J, Wiggins L, Christensen DL, Maenner MJ, Daniels J, Warren Z, et al (2018) Prevalence of autism spectrum disorder among children aged 8 Years - Autism and developmental disabilities monitoring network, 11 Sites, United States, 2014. MMWR Surveill Summ. https://doi.org/10.15585/mmwr.ss6706a1
Doyle CA, McDougle CJ (2012) Pharmacologic treatments for the behavioral symptoms associated with autism spectrum disorders across the lifespan. Dialogues Clin Neurosci. 14:263–279
Brentani H, de Paula CS, Bordini D, Rolim D, Sato F, Portolese J, et al (2013) Autism spectrum disorders: an overview on diagnosis and treatment. Rev Bras Psiquiatr. https://doi.org/10.1590/1516-4446-2013-S104
Mody M, Belliveau JW (2012) Speech and language impairments in autism: insights from behavior and neuroimaging. Am Chin J Med Sci. https://doi.org/10.7156/v5i3p157
Holt R, Barnby G, Maestrini E, Bacchelli E, Brocklebank D, Sousa I, et al (2010) Linkage and candidate gene studies of autism spectrum disorders in European populations. Eur J Hum Genet. https://doi.org/10.1038/ejhg.2010.69
Grove J, Ripke S, Als TD, Mattheisen M, Walters RK, Won H, et al (2019) Identification of common genetic risk variants for autism spectrum disorder. Nat Genet. https://doi.org/10.1038/s41588-019-0344-8
McBride KL, Varga EA, Pastore MT, Prior TW, Manickam K, Atkin JF, et al (2010) Confirmation study of PTEN mutations among individuals with autism or developmental delays/mental retardation and macrocephaly. Autism Res. https://doi.org/10.1002/aur.132
Vaishnavi V, Manikandan M, Munirajan AK (2014) Mining the 3’UTR of Autism-implicated genes for SNPs perturbing MicroRNA regulation. Genomics Proteomics Bioinforma. https://doi.org/10.1016/j.gpb.2014.01.003
Alonso-Gonzalez A, Rodriguez-Fontenla C, Carracedo A (2018) De novo mutations (DNMs) in autism spectrum disorder (ASD): pathway and network analysis. Front Genet. https://doi.org/10.3389/fgene.2018.00406
Al-Mubarak B, Abouelhoda M, Omar A, Aldhalaan H, Aldosari M, Nester M, et al (2017) Whole exome sequencing reveals inherited and de novo variants in autism spectrum disorder: A trio study from Saudi families. Sci Rep. https://doi.org/10.1038/s41598-017-06033-1
Sanders SJ (2019) Next-generation sequencing in autism spectrum disorder. Cold Spring Harb Perspect Med. https://doi.org/10.1101/cshperspect.a026872
Rosain J, Oleaga-Quintas C, Deswarte C, Verdin H, Marot S, Syridou G, et al (2018) Avarietyofalu-mediated copy number variations can underlie il-12rβ1 deficiency. J Clin Immunol. https://doi.org/10.1007/s10875-018-0527-6
D’Gama AM, Pochareddy S, Li M, Jamuar SS, Reiff RE, Lam ATN, et al (2015) Targeted DNA sequencing from autism spectrum disorder brains implicates multiple genetic mechanisms. Neuron. https://doi.org/10.1016/j.neuron.2015.11.009
Jiang YH, Wang Y, Xiu X, Choy KW, Pursley AN, Cheung SW (2014) Genetic diagnosis of autism spectrum disorders: the opportunity and challenge in the genomics era. Crit Rev Clin Lab Sci. https://doi.org/10.3109/10408363.2014.910747
Dang VT, Kassahn KS, Marcos AE, Ragan MA (2008) Identification of human haploinsufficient genes and their genomic proximity to segmental duplications. Eur J Hum Genet. https://doi.org/10.1038/ejhg.2008.111
Ji X, Kember RL, Brown CD, Bućan M (2016) Increased burden of deleterious variants in essential genes in autism spectrum disorder. Proc Natl Acad Sci USA. https://doi.org/10.1073/pnas.1613195113
Abuín JM, Pichel JC, Pena TF, Amigo J (2016) SparkBWA: speeding up the alignment of high-throughput DNA sequencing data. PLoS One. https://doi.org/10.1371/journal.pone.0155461
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods. https://doi.org/10.1038/nmeth.1923
Griswold AJ, Dueker ND, Van Booven D, Rantus JA, Jaworski JM, Slifer SH, et al (2015) Targeted massively parallel sequencing of autism spectrum disorder-associated genes in a case control cohort reveals rare loss-of-function risk variants. Mol Autism. https://doi.org/10.1186/s13229-015-0034-z
Huang N, Lee I, Marcotte EM, Hurles ME (2010) Characterising and predicting haploinsufficiency in the human genome. PLoS Genet. https://doi.org/10.1371/journal.pgen.1001154
Fuller ZL, Berg JJ, Mostafavi H, Sella G, Przeworski M (2019) Measuring intolerance to mutation in human genetics. Nat Genet. https://doi.org/10.1038/s41588-019-0383-1
Vishweswaraiah S, Veerappa AM, Mahesh PA, Jahromi SR, Ramachandra NB (2015) Copy number variation burden on asthma subgenome in normal cohorts identifies susceptibility markers. Allergy, Asthma Immunol Res. https://doi.org/10.4168/aair.2015.7.3.265
Murthy MN, Veerappa AM, Seshachalam KB, Ramachandra NB (2016) High-resolution arrays reveal burden of copy number variations on Parkinson disease genes associated with increased disease risk in random cohorts. Neurol Res. https://doi.org/10.1080/01616412.2016.1204105
Suresh RV, Lingaiah K, Veerappa AM, Ramachandra NB (2017) Identifying the risk of producing aneuploids using meiotic recombination genes as biomarkers: a copy number variation approach. Indian J Med Res. https://doi.org/10.4103/ijmr.IJMR_965_14
Gholipoorfeshkecheh R, Agarwala S, Krishnappa S, Savitha MR, Narayanappa D, Ramachandra NB (2020) Variants in HEY genes manifest in ventricular septal defects of congenital heart disease. Gene Reports. https://doi.org/10.1016/j.genrep.2020.100613
Quan L, Lv Q, Zhang YSTRUM (2016) Structure-based prediction of protein stability changes upon single-point mutation. Bioinformatics. https://doi.org/10.1093/bioinformatics/btw361
Marsden RL, McGuffin LJ, Jones DT (2009) Rapid protein domain assignment from amino acid sequence using predicted secondary structure. Protein Sci. https://doi.org/10.1110/ps.0209902
Peñagarikano O, Geschwind DH (2012) What does CNTNAP2 reveal about autism spectrum disorder? Trends Mol Med. https://doi.org/10.1016/j.molmed.2012.01.003
Koemans TS, Kleefstra T, Chubak MC, Stone MH, Reijnders MRF, de Munnik S, et al (2017) Functional convergence of histone methyltransferases EHMT1 and KMT2C involved in intellectual disability and autism spectrum disorder. PLoS Genet. https://doi.org/10.1371/journal.pgen.1006864
Feliciano P, Zhou X, Astrovskaya I, Turner TN, Wang T, Brueggeman L, et al (2019) Exome sequencing of 457 autism families recruited online provides evidence for autism risk genes. npj Genom Med. https://doi.org/10.1038/s41525-019-0093-8
Schaaf CP, Sabo A, Sakai Y, Crosby J, Muzny D, Hawes A, et al (2011) Oligogenic heterozygosity in individuals with high-functioning autism spectrum disorders. Hum Mol Genet. https://doi.org/10.1093/hmg/ddr243
Rossi M, El-Khechen D, Black MH, Farwell Hagman KD, Tang S, Powis Z (2017) Outcomes of Diagnostic Exome Sequencing in Patients With Diagnosed or Suspected Autism Spectrum Disorders. Pediatr Neurol. https://doi.org/10.1016/j.pediatrneurol.2017.01.033
Satterstrom FK, Kosmicki JA, Wang J, Breen MS, de Rubeis S, An JY, et al (2020) Large-Scale Exome Sequencing Study Implicates Both Developmental and Functional Changes in the Neurobiology of Autism. Cell. https://doi.org/10.1016/j.cell.2019.12.036
Yuen RKC, Merico D, Bookman M, Howe JL, Thiruvahindrapuram B (2017) Patel Rv, et al. Whole genome sequencing resource identifies 18 new candidate genes for autism spectrum disorder. Nat Neurosci. https://doi.org/10.1038/nn.4524
de Rubeis S, He X, Goldberg AP, Poultney CS, Samocha K, Cicek AE, et al (2014) Synaptic, transcriptional and chromatin genes disrupted in autism. Nature. https://doi.org/10.1038/nature13772
Iossifov I, O’Roak BJ, Sanders SJ, Ronemus M, Krumm N, Levy D, et al (2014) The contribution of de novo coding mutations to autism spectrum disorder. Nature. https://doi.org/10.1038/nature13908
Krumm N, Turner TN, Baker C, Vives L, Mohajeri K, Witherspoon K, et al (2015) Excess of rare, inherited truncating mutations in autism. Nat Genet. https://doi.org/10.1038/ng.3303
Wang T, Guo H, Xiong B, Stessman HAF, Wu H, Coe BP, et al (2016) De novo genic mutations among a Chinese autism spectrum disorder cohort. Nat Commun. https://doi.org/10.1038/ncomms13316
Li TD, Guo CR, Lan LY, Ke LW, Fang ZY, Kai LJ, et al (2019) The critical role of ASD-related gene CNTNAP3 in regulating synaptic development and social behavior in mice. Neurobiol Dis. https://doi.org/10.1016/j.nbd.2019.104486
Turner TN, Coe BP, Dickel DE, Hoekzema K, Nelson BJ, Zody MC, et al (2017) Genomic Patterns of De Novo Mutation in Simplex Autism. Cell. https://doi.org/10.1016/j.cell.2017.08.047
Klassen T, Davis C, Goldman A, Burgess D, Chen T, Wheeler D, et al (2011) Exome sequencing of ion channel genes reveals complex profiles confounding personal risk assessment in epilepsy. Cell. https://doi.org/10.1016/j.cell.2011.05.025
Torrico B, Shaw AD, Mosca R, Vivó-Luque N, Hervás A, Fernàndez-Castillo N, et al (2019) Truncating variant burden in high-functioning autism and pleiotropic effects of LRP1 across psychiatric phenotypes. J Psychiatry Neurosci. https://doi.org/10.1503/jpn.180184
Sener EF (2014) Association of Copy Number Variations in Autism Spectrum Disorders: A Systematic Review. Chin J Biol. https://doi.org/10.1155/2014/713109
Spencer DH, Tyagi M, Vallania F, Bredemeyer AJ, Pfeifer JD, Mitra RD, et al (2014) Performance of common analysis methods for detecting low-frequency single nucleotide variants in targeted next-generation sequence data. J Mol Diagn. https://doi.org/10.1016/j.jmoldx.2013.09.003
Chiara M, Pavesi G (2017) Evaluation of quality assessment protocols for high throughput genome resequencing data. Front Genet. https://doi.org/10.3389/fgene.2017.00094
Abrahams BS, Arking DE, Campbell DB, Mefford HC, Morrow EM, Weiss LA, et al (2013) SFARI Gene 2.0: a community-driven knowledgebase for the autism spectrum disorders (ASDs). Mol Autism. https://doi.org/10.1186/2040-2392-4-36
Sestan N, State MW (2018) Lost in translation: traversing the complex path from genomics to therapeutics in autism spectrum disorder. Neuron. https://doi.org/10.1016/j.neuron.2018.10.015
Rylaarsdam L, Guemez-Gamboa A (2019) Genetic causes and modifiers of autism spectrum disorder. Front Cell Neurosci. https://doi.org/10.3389/fncel.2019.00385
Lavery WJ, Barski A, Wiley S, Schorry EK, Lindsley AW (2020) KMT2C/D COMPASS complex-associated diseases [KCDCOM-ADs]: An emerging class of congenital regulopathies. Clin Epigenetics. https://doi.org/10.1186/s13148-019-0802-2
Yong-Hui J, Ehlers MD (2013) Modeling autism by SHANK gene mutations in mice. Neuron. https://doi.org/10.1016/j.neuron.2013.03.016
Uchino S, Waga C (2015) Novel therapeutic approach for autism spectrum disorder: focus on SHANK3. Curr Neuropharmacol. https://doi.org/10.2174/1570159x13666151029105547
Guang S, Pang N, Deng X, Yang L, He F, Wu L et al (2018) Synaptopathology involved in autism spectrum disorder. Front Cell Neurosci. https://doi.org/10.3389/fncel.2018.00470
Lu AT-H, Dai X, Martinez-Agosto JA, Cantor RM (2012) Support for calcium channel gene defects in autism spectrum disorders. Mol Autism. https://doi.org/10.1186/2040-2392-3-18
Toma C, Pierce KD, Shaw AD, Heath A, Mitchell PB, Schofield PR et al (2018) Comprehensive cross-disorder analyses of CNTNAP2 suggest it is unlikely to be a primary risk gene for psychiatric disorders. PLoS Genet. https://doi.org/10.1371/journal.pgen.1007535
Sampath S, Bhat S, Gupta S, O’Connor A, West AB, Arking DE et al (2013) Defining the contribution of CNTNAP2 to autism susceptibility. PLoS One. https://doi.org/10.1371/journal.pone.0077906
Brueggeman L, Koomar T, Michaelson JJ (2020) Author correction: forecasting risk gene discovery in autism with machine learning and genome-scale data. (Scientific Reports, (2020), 10, 1, (4569), 10.1038/s41598-020-61288-5). Sci Rep. https://doi.org/10.1038/s41598-020-77832-2
Yang R, Walder-Christensen KK, Kim N, Wu D, Lorenzo DN, Badea A et al (2019) ANK2 autism mutation targeting giant ankyrin-B promotes axon branching and ectopic connectivity. Proc Natl Acad Sci U S A. https://doi.org/10.1073/pnas.1904348116
Cheroni C, Caporale N, Testa G (2020) Autism spectrum disorder at the crossroad between genes and environment: contributions, convergences, and interactions in ASD developmental pathophysiology. Mol Autism. https://doi.org/10.1186/s13229-020-00370-1
Canali G, Goutebroze L (2018) CNTNAP2 heterozygous missense variants: risk factors for autism spectrum disorder and/or other pathologies? J Exp Neurosci. https://doi.org/10.1177/1179069518809666
Rodenas-Cuadrado P, Ho J, Vernes SC (2014) Shining a light on CNTNAP2: complex functions to complex disorders. Eur J Hum Genet. https://doi.org/10.1038/ejhg.2013.100
Aylward A, Cai Y, Lee A, Blue E, Rabinowitz D, Haddad J (2016) Using whole exome sequencing to identify candidate genes with rare variants in nonsyndromic cleft lip and palate. Genet Epidemiol. https://doi.org/10.1002/gepi.21972
Redler RL, Das J, Diaz JR, Dokholyan NV (2016) Protein destabilization as a common factor in diverse inherited disorders. J Mol Evol. https://doi.org/10.1007/s00239-015-9717-5
Russo AJ (2014) Increased epidermal growth factor receptor (EGFR) associated with hepatocyte growth factor (HGF) and symptom severity in children with autism spectrum disorders (ASDs). J Cent Nerv Syst Dis. https://doi.org/10.4137/jcnsd.s13767
Lou CH, Shao A, Shum EY, Espinoza JL, Huang L, Karam R et al (2014) Posttranscriptional control of the stem cell and neurogenic programs by the nonsense-mediated RNA decay pathway. Cell Rep. https://doi.org/10.1016/j.celrep.2014.01.028
Vaishnavi V, Manikandan M, Tiwary BK, Munirajan AK (2013) Insights on the functional impact of microRNAs present in autism-associated copy number variants. PLoS One. https://doi.org/10.1371/journal.pone.0056781
Poot M (2015) Connecting the CNTNAP2 networks with neurodevelopmental disorders. Mol Syndromol. https://doi.org/10.1159/000371594
Thiltgen G, Goldstein RA (2012) Assessing predictors of changes in protein stability upon mutation using self-consistency. PLoS One. https://doi.org/10.1371/journal.pone.0046084
Goldman N, Thorne JL, Jones DT (1998) Assessing the impact of secondary structure and solvent accessibility on protein evolution. Genetics. 149:445–458
Lins L, Thomas A, Brasseur R (2003) Analysis of accessible surface of residues in proteins. Protein Sci. https://doi.org/10.1110/ps.0304803
Receveur-Brechot V, Durand D (2012) How random are intrinsically disordered proteins? A small angle scattering perspective. Curr Protein Pept Sci. https://doi.org/10.2174/138920312799277901
Macossay-Castillo M, Kosol S, Tompa P, Pancsa R (2014) Synonymous constraint elements show a tendency to encode intrinsically disordered protein segments. PLoS Comput Biol. https://doi.org/10.1371/journal.pcbi.1003607
Chen P, Liu C, Burge L, Li J, Mohammad M, Southerland W et al (2010) DomSVR: Domain boundary prediction with support vector regression from sequence information alone. Amino Acids. https://doi.org/10.1007/s00726-010-0506-6
Poot M (2017) Intragenic CNTNAP2 deletions: a bridge too far? Mol Syndromol. https://doi.org/10.1159/000456021
Vernes SC, Newbury DF, Abrahams BS, Winchester L, Nicod J, Groszer M et al (2008) A functional genetic link between distinct developmental language disorders. N Engl J Med. https://doi.org/10.1056/NEJMoa0802828
Nguyen RL, Medvedeva YV, Ayyagari TE, Schmunk G, Gargus JJ (2018) Intracellular calcium dysregulation in autism spectrum disorder: an analysis of converging organelle signaling pathways. Biochim Biophys Acta, Mol Cell Res. https://doi.org/10.1016/j.bbamcr.2018.08.003
Acknowledgements
We thank the subjects and their families for participating in this study; the University of Mysore for their help and encouragement; the Department of Studies in Genetics and Genomics, University of Mysore for providing facility to conduct this work; and also the research colleges from our laboratory, Department of Studies in Genetics and Genomics, University of Mysore for their support.
Funding
(1) Department of Science and Technology—Health Science, Government of India, New Delhi (DST/INSPIRE/IF160260) and (2) University Grants Commission-Major Research Project (MRP-MAJOR-GENE- 2013-19809) helped in the funding needed for the study and validations.
Author information
Authors and Affiliations
Contributions
SA has made substantial contributions to the conception and design, and analysis and interpretation of data and is involved in drafting the manuscript and reviewing the manuscript critically. NBR has made substantial contributions to the conception and design, revised the manuscript critically for important intellectual content, and agreed to be accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Written consent was obtained from all participants involved in this study, and the Institutional Human Ethical Committee (IHEC No.128 Ph.D/2016–17) approved the consent procedure
Consent for publication
Consent has been taken from all participants, the data provider, and the authors for the said publication
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Supplementary Figure 1
. Protein-protein interactions of CNTNAP2 with autism-relevant NrCAM, ANK2, and CNTN4 proteins present in sample dataset under study along with comparative gene expression levels
Additional file 2: Supplementary Figure 2
. Gene mutational landscape with SFARI gene scoring for the sample cohort under investigation reveals autism genes with categorical gene scores
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Agarwala, S., Ramachandra, N.B. Role of CNTNAP2 in autism manifestation outlines the regulation of signaling between neurons at the synapse. Egypt J Med Hum Genet 22, 22 (2021). https://doi.org/10.1186/s43042-021-00138-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1186/s43042-021-00138-z